| This post originally appeared on the Rittman Mead blog. |
In the previous articles (here, and here) I gave the background to a project we did for a client, exploring the benefits of Spark-based ETL processing running on Amazon’s Elastic Map Reduce (EMR) Hadoop platform. The proof of concept we ran was on a very simple requirement, taking inbound files from a third party, joining to them to some reference data, and then making the result available for analysis.
I showed here how I built up the prototype PySpark code on my local machine, using Docker to quickly and easily make available the full development environment needed.
Now it’s time to get it running on a proper Hadoop platform. Since the client we were working with already have a big presence on Amazon Web Services (AWS), using Amazon’s Hadoop platform made sense. Amazon’s Elastic Map Reduce, commonly known as EMR, is a fully configured Hadoop cluster. You can specify the size of the cluster and vary it as you want (hence, "Elastic"). One of the very powerful features of it is that being a cloud service, you can provision it on demand, run your workload, and then shut it down. Instead of having a rack of physical servers running your Hadoop platform, you can instead spin up EMR whenever you want to do some processing - to a size appropriate to the processing required - and only pay for the processing time that you need.
Moving my locally-developed PySpark code to run on EMR should be easy, since they’re both running Spark. Should be easy, right? Well, this is where it gets - as we say in the trade - "interesting". Part of my challenges were down to the learning curve in being new to this set of technology. However, others I would point to more as being examples of where the brave new world of Big Data tooling becomes less an exercise in exciting endless possibilities and more stubbornly Googling errors due to JAR clashes and software version mismatches…
Provisioning EMR 🔗
Whilst it’s possible to make the entire execution of the PySpark job automated (including the provisioning of the EMR cluster itself), to start with I wanted to run it manually to check each step along the way.
To create an EMR cluster simply login to the EMR console and click Create
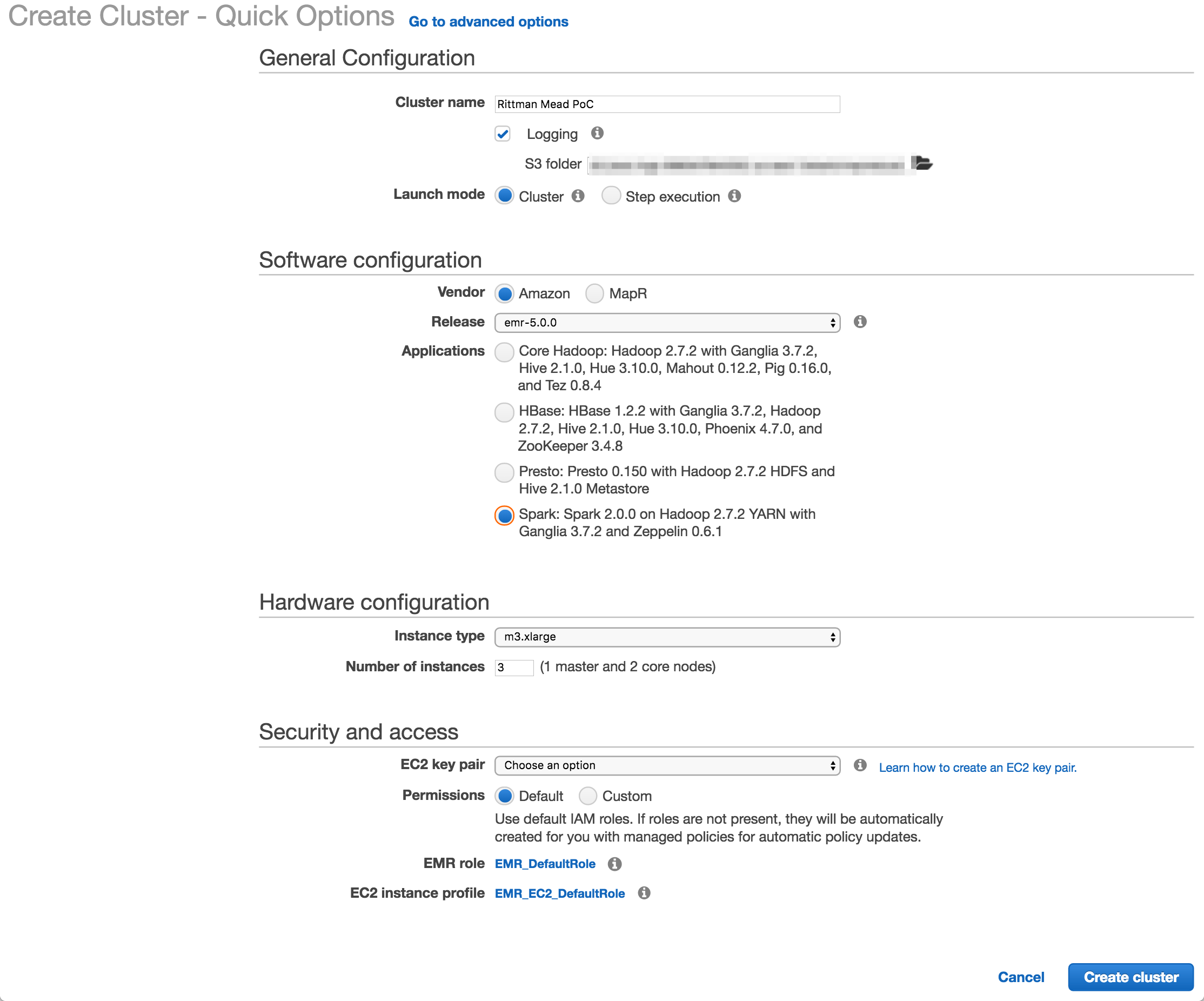
I used Amazon’s EMR distribution, configured for Spark. You can also deploy a MapR-based hadoop platform, and use the Advanced tab to pick and mix the applications to deploy (such as Spark, Presto, etc).
The number and size of the nodes is configured here (I used the default, 3 machines of m3.xlarge spec), as is the SSH key. The latter is very important to get right, otherwise you won’t be able to connect to your cluster over SSH.
Once you click Create cluster Amazon automagically provisions the underlying EC2 servers, and deploys and configures the software and Hadoop clustering across them. Anyone who’s set up a Hadoop cluster will know that literally a one-click deploy of a cluster is a big deal!
If you’re going to be connecting to the EMR cluster from your local machine you’ll want to modify the security group assigned to it once provisioned and enable access to the necessary ports (e.g. for SSH) from your local IP.
Deploying the code 🔗
I developed the ETL code in Jupyter Notebooks, from where it’s possible to export it to a variety of formats - including .py Python script. All the comment blocks from the Notebook are carried across as inline code comments.
To transfer the Python code to the EMR cluster master node I initially used scp, simply out of habit. But, a much more appropriate solution soon presented itself - S3! Not only is this a handy way of moving data around, but it comes into its own when we look at automating the EMR execution later on.
To upload a file to S3 you can use the S3 web interface, or a tool such as Cyberduck. Better, if you like the command line as I do, is the AWS CLI tools. Once installed, you can run this from your local machine:
aws s3 cp Acme.py s3://foobar-bucket/code/Acme.pyYou’ll see that the syntax is pretty much the same as the Linux cp comand, specifying source and then destination. You can do a vast amount of AWS work from this command line tool - including provisioning EMR clusters, as we’ll see shortly.
So with the code up on S3, I then SSH’d to the EMR master node (as the hadoop user, not ec2-user), and transfered it locally. One of the nice things about EMR is that it comes with your AWS security automagically configred. Whereas on my local machine I need to configure my AWS credentials in order to use any of the aws commands, on EMR the credentials are there already.
aws s3 cp s3://foobar-bucket/code/Acme.py ~This copied the Python code down into the home folder of the hadoop user.
Running the code - manually 🔗
To invoke the code, simply run:
spark-submit Acme.pyA very useful thing to use, if you aren’t already, is GNU screen (or tmux, if that’s your thing). GNU screen is installed by default on EMR (as it is on many modern Linux distros nowadays). Screen does lots of cool things, but of particular relevance here is it lets you close your SSH connection whilst keeping your session on the server open and running. You can then reconnect at a later time back to it, and pick up where you left off. Whilst you’re disconnected, your session is still running and the work still being processed.
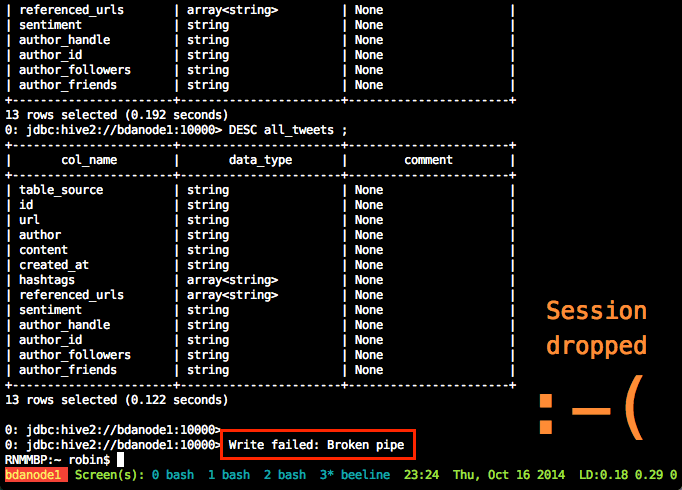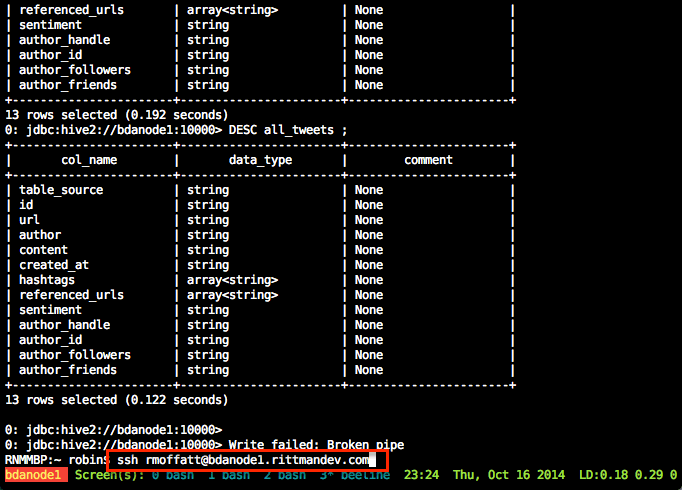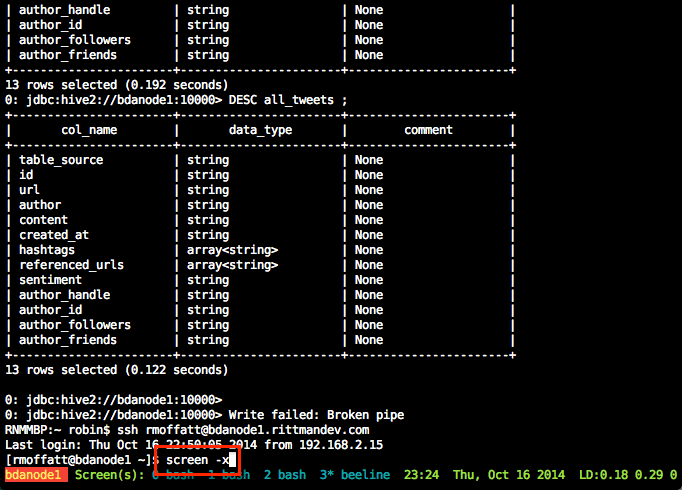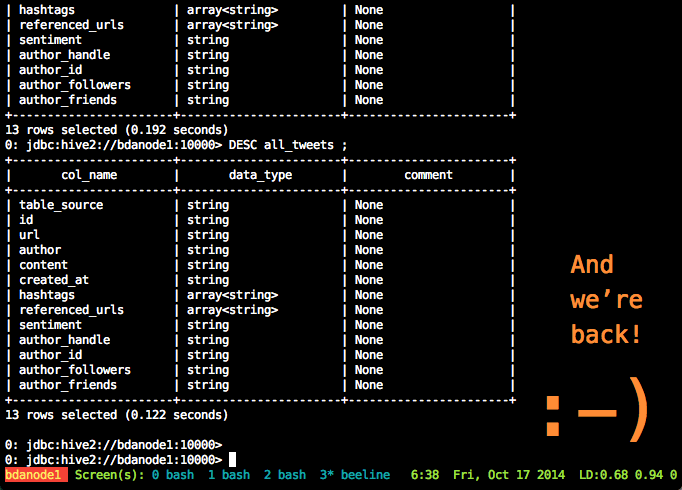
From the Spark console you can monitor the execution of the job running, as well as digging into the details of how it undertakes the work. See the EMR cluster home page on AWS for the Spark console URL
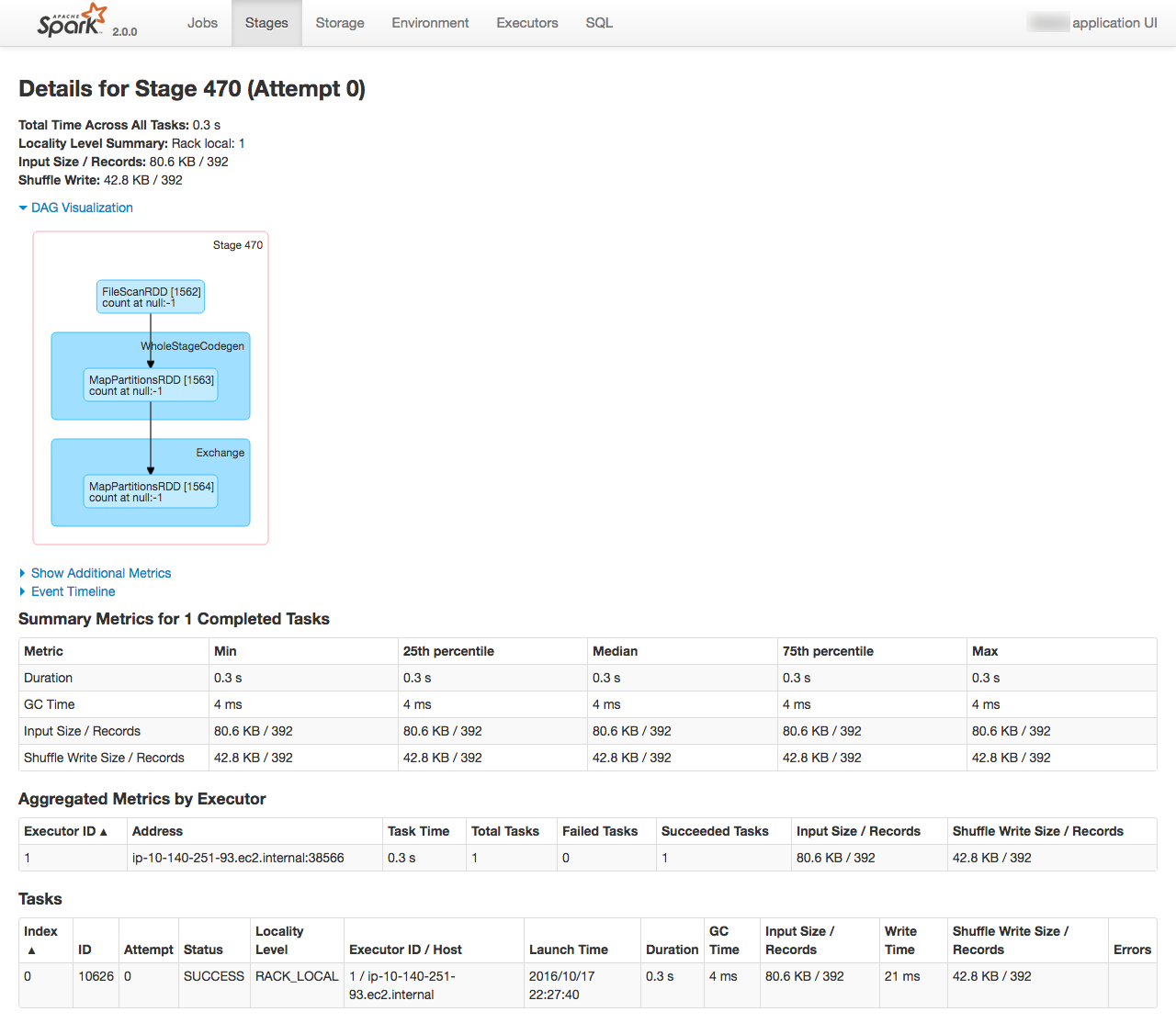
| This post originally appeared on the Rittman Mead blog. |
